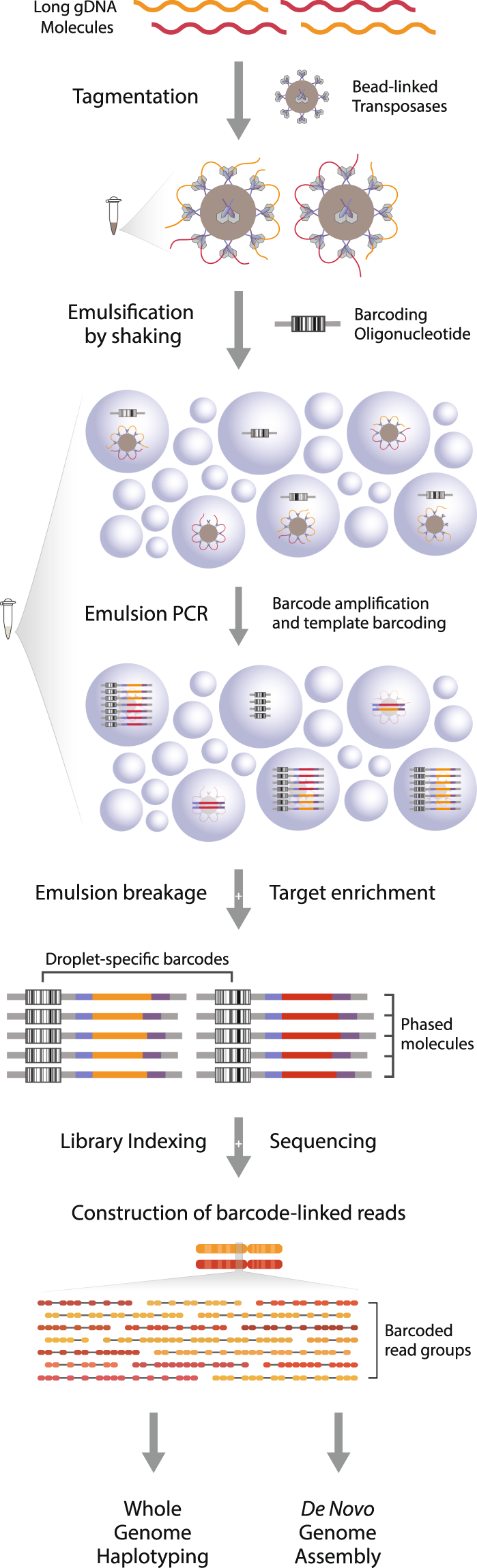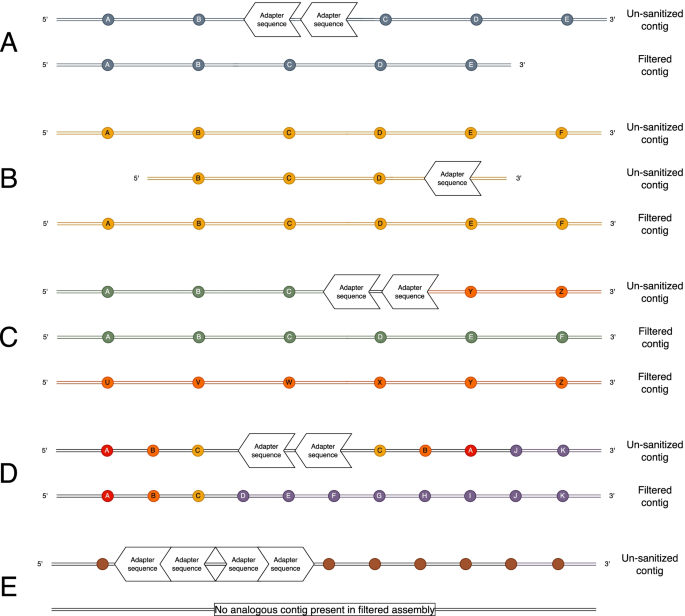
HiFiAdapterFilt, a memory efficient read processing pipeline, prevents occurrence of adapter sequence in PacBio HiFi reads and their negative impacts on genome assembly | BMC Genomics | Full Text
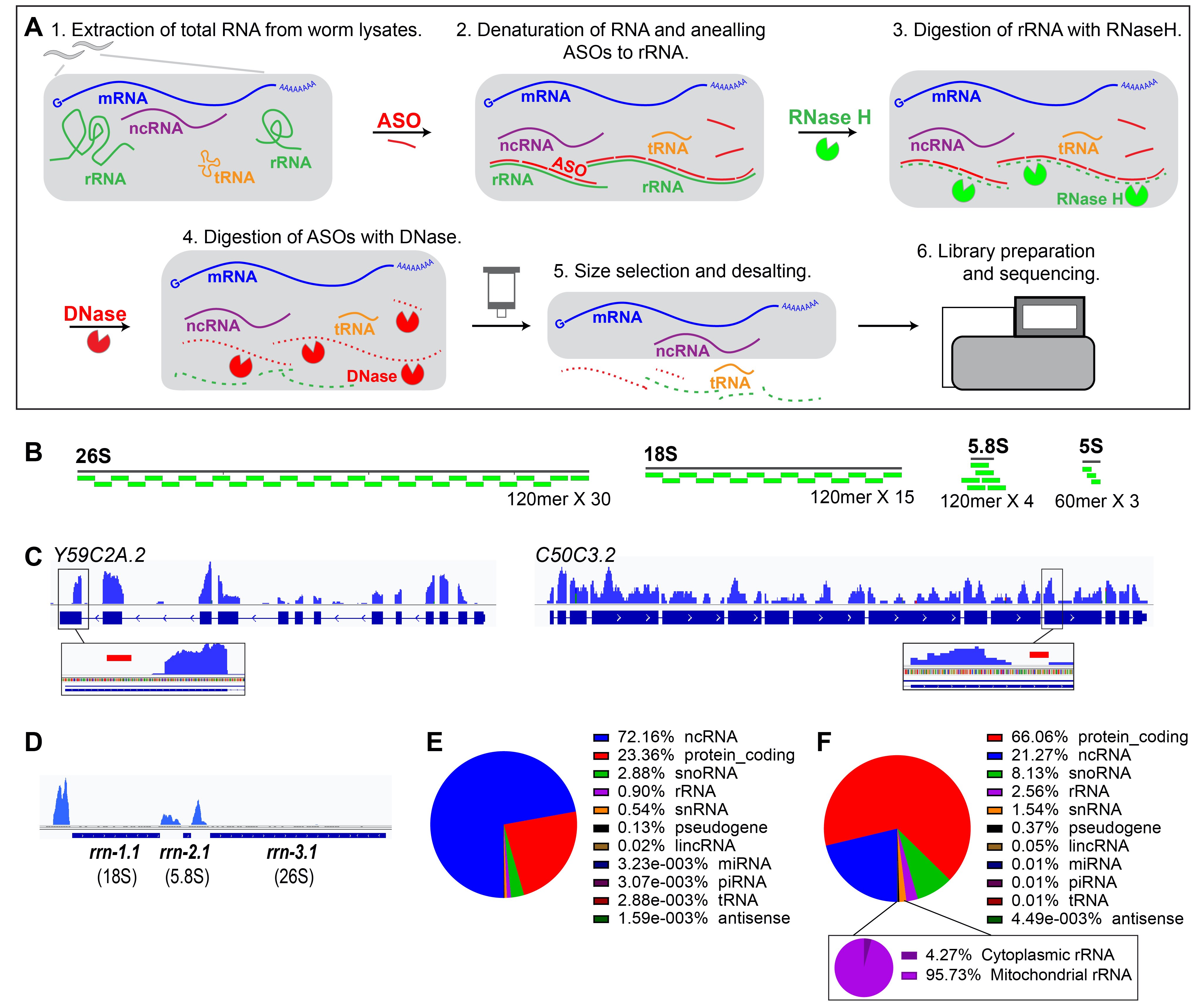
RNA-seq with RNase H-based ribosomal RNA depletion specifically designed for C. elegans | microPublication

Aptardi predicts polyadenylation sites in sample-specific transcriptomes using high-throughput RNA sequencing and DNA sequence | Nature Communications
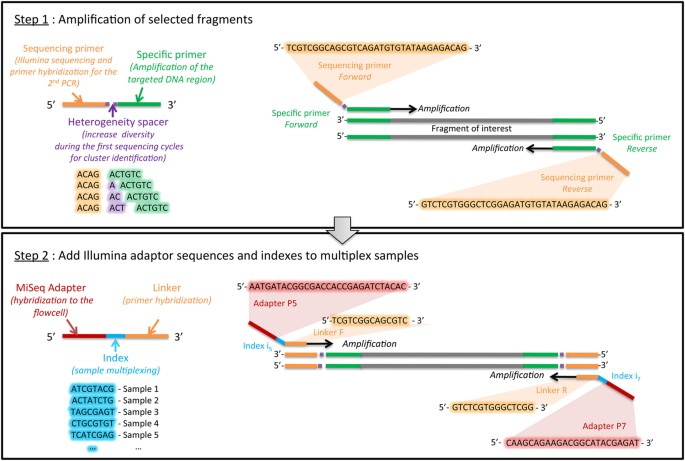
High-throughput sequencing of multiple amplicons for barcoding and integrative taxonomy | Scientific Reports
Comparison of tools cutting primer sequences from reads with tolerance... | Download Scientific Diagram

FASTAptameR 2.0: A web tool for combinatorial sequence selections: Molecular Therapy - Nucleic Acids
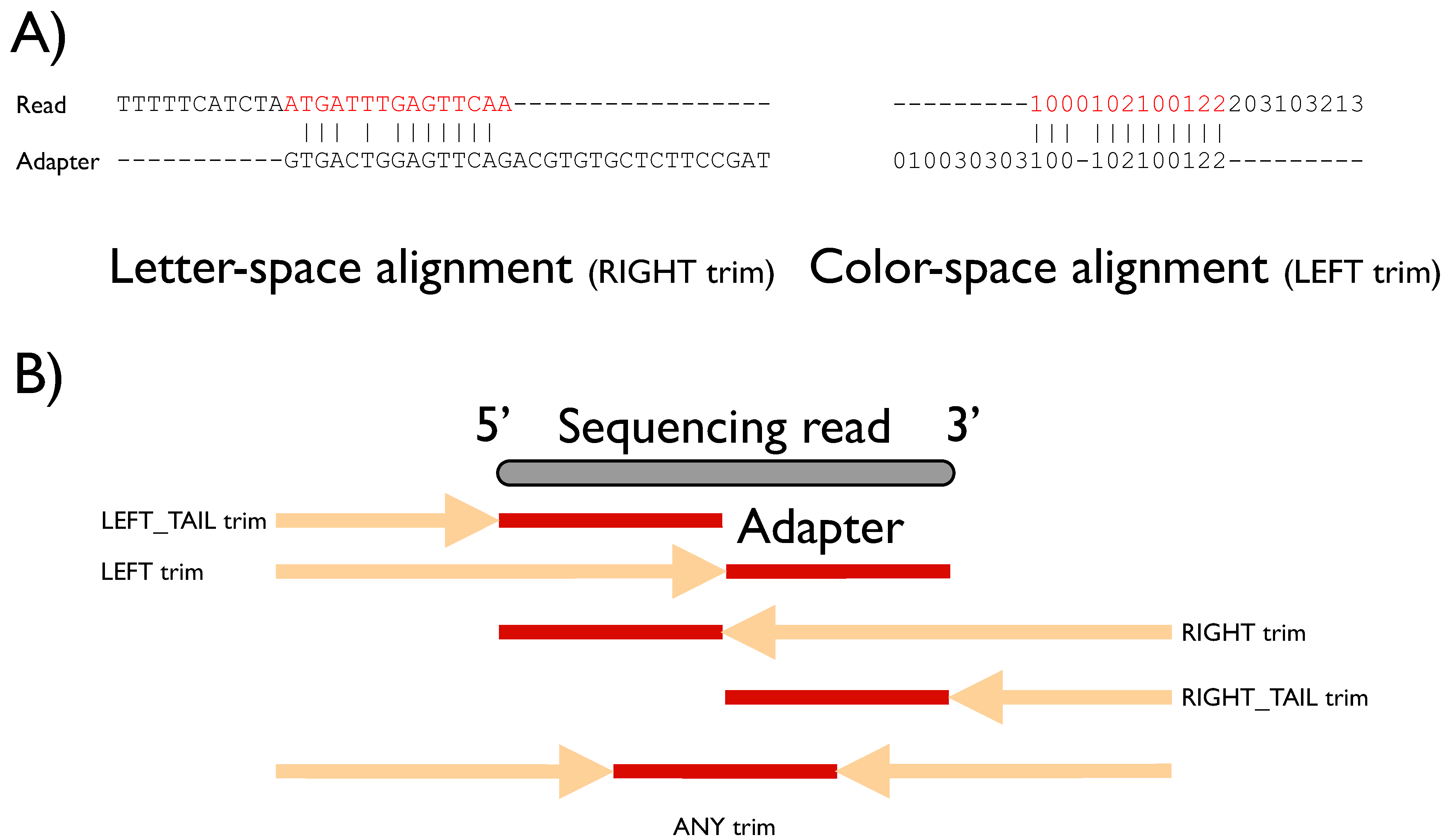
Biology | Free Full-Text | FLEXBAR—Flexible Barcode and Adapter Processing for Next-Generation Sequencing Platforms

FASTAptamer: A Bioinformatic Toolkit for High-throughput Sequence Analysis of Combinatorial Selections: Molecular Therapy - Nucleic Acids

Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports

Kraken: A set of tools for quality control and analysis of high-throughput sequence data☆ | Semantic Scholar
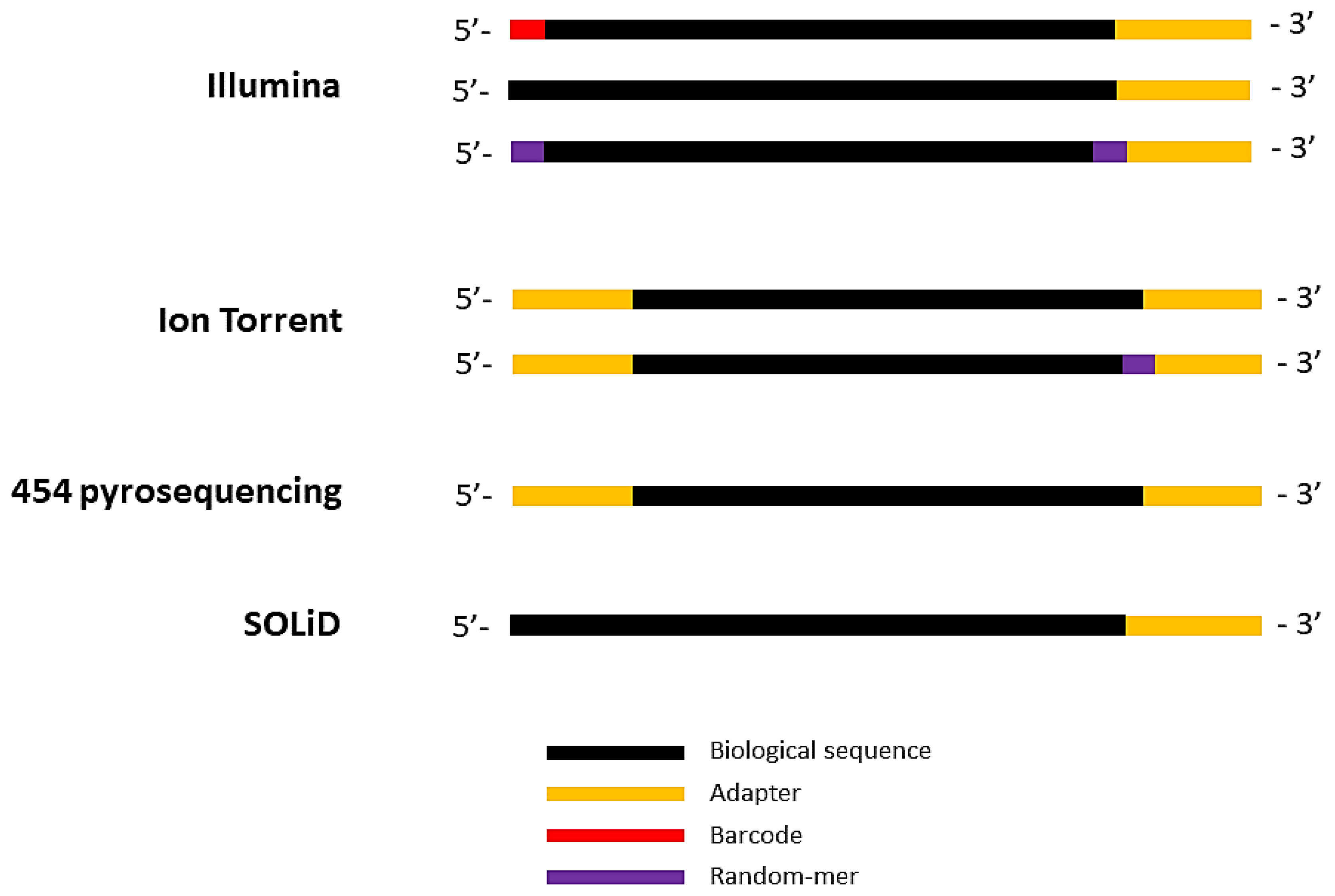
Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data

cutPrimers: A New Tool for Accurate Cutting of Primers from Reads of Targeted Next Generation Sequencing | Journal of Computational Biology
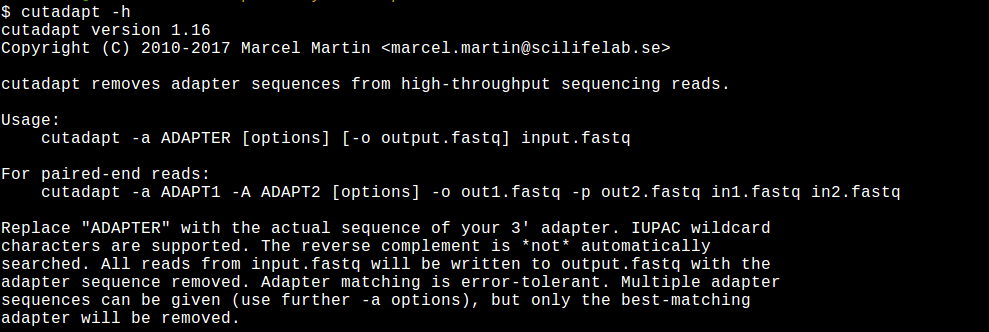
Cutadapt remoYes adapter sequences from high-throughput sequencing reads Introduction Implementation Features